Please wait while we process your payment
If you don't see it, please check your spam folder. Sometimes it can end up there.
If you don't see it, please check your spam folder. Sometimes it can end up there.
Please wait while we process your payment
Get instant, ad-free access to our grade-boosting study tools with a 7-day free trial!
Learn more



Create Account
This site is protected by reCAPTCHA and the Google Privacy Policy and Terms of Service apply.
Log into your PLUS account
Create Account
Select Plan
Payment Info
Start 7-Day Free Trial!
Select Your Plan
Monthly
$5.99
/month + tax
Annual
$29.99
/year + taxAnnual
2-49 accounts
$22.49/year + tax
50-99 accounts
$20.99/year + tax
Select Quantity
Price per seat
$29.99 $--.--
Subtotal
$-.--
Want 100 or more? Request a customized plan
Monthly
$5.99
/month + taxYou could save over 50%
by choosing an Annual Plan!

Annual
$29.99
/year + taxSAVE OVER 50%
compared to the monthly price!
| Focused-studying | ||
| PLUS Study Tools | ||
| AP® Test Prep PLUS | ||
| My PLUS Activity | ||
Annual
$22.49/month + tax
Save 25%
on 2-49 accounts
Annual
$20.99/month + tax
Save 30%
on 50-99 accounts
| Focused-studying | ||
| PLUS Study Tools | ||
| AP® Test Prep PLUS | ||
| My PLUS Activity | ||
Testimonials from SparkNotes Customers
No Fear provides access to Shakespeare for students who normally couldn’t (or wouldn’t) read his plays. It’s also a very useful tool when trying to explain Shakespeare’s wordplay!
Erika M.
I tutor high school students in a variety of subjects. Having access to the literature translations helps me to stay informed about the various assignments. Your summaries and translations are invaluable.
Kathy B.
Teaching Shakespeare to today's generation can be challenging. No Fear helps a ton with understanding the crux of the text.
Kay H.
Testimonials from SparkNotes Customers
No Fear provides access to Shakespeare for students who normally couldn’t (or wouldn’t) read his plays. It’s also a very useful tool when trying to explain Shakespeare’s wordplay!
Erika M.
I tutor high school students in a variety of subjects. Having access to the literature translations helps me to stay informed about the various assignments. Your summaries and translations are invaluable.
Kathy B.
Teaching Shakespeare to today's generation can be challenging. No Fear helps a ton with understanding the crux of the text.
Kay H.
Create Account
Select Plan
Payment Info
Start 7-Day Free Trial!
Payment Information
You will only be charged after the completion of the 7-day free trial.
If you cancel your account before the free trial is over, you will not be charged.
You will only be charged after the completion of the 7-day free trial. If you cancel your account before the free trial is over, you will not be charged.
Order Summary
Annual
7-day Free Trial
SparkNotes PLUS
$29.99 / year
Annual
Quantity
51
PLUS Group Discount
$29.99 $29.99 / seat
Tax
$0.00
SPARK25
-$1.25
25% Off
Total billed on Nov 7, 2024 after 7-day free trail
$29.99
Total billed
$0.00
Due Today
$0.00
Promo code
This is not a valid promo code
Card Details
By placing your order you agree to our terms of service and privacy policy.
By saving your payment information you allow SparkNotes to charge you for future payments in accordance with their terms.
Powered by stripe
Legal
Google pay.......



Thank You!
Your group members can use the joining link below to redeem their membership. They will be prompted to log into an existing account or to create a new account. All members under 16 will be required to obtain a parent's consent sent via link in an email.Your Child’s Free Trial Starts Now!
Thank you for completing the sign-up process. Your child’s SparkNotes PLUS login credentials are [email] and the associated password. If you have any questions, please visit our help center.Your Free Trial Starts Now!
Please wait while we process your payment

Sorry, you must enter a valid email address
By entering an email, you agree to our privacy policy.
Please wait while we process your payment

Sorry, you must enter a valid email address
By entering an email, you agree to our privacy policy.
Please wait while we process your payment

Your PLUS subscription has expired
Please wait while we process your payment
Please wait while we process your payment
Month
Day
Year
Please read our terms and privacy policy
Please wait while we process your payment

DNA Proof-Reading and Repair
The low overall rate of mutation during DNA replication (1 base pair change in one billion base pairs per replication cycle) does not reflect the true number of errors that take place during the replication process. The number is kept so low by a proof-reading system that checks newly synthesized DNA for errors and corrects them when they are found. Errors in DNA replication can take different forms, but usually revolve around the addition of a nucleotide with the incorrect base, meaning the pairing between the parent and daughter strand bases is not complementary. The addition of an incorrect base can take place by a process called tautomerization. A tautomer of a base group is a slight rearrangement of its electrons that allows for different bonding patterns between bases. This can lead to the incorrect pairing of C with A instead of G, for example.
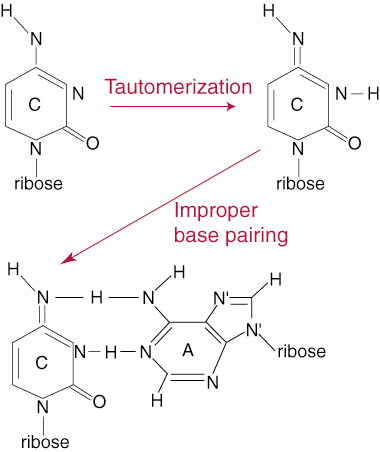
DNA retains its high level of accuracy is with its proof-reading function.
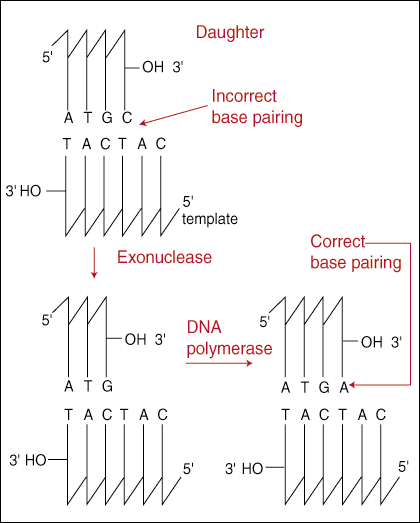
The 3' to 5' proof-reading exonuclease works by scanning along directly behind as the DNA polymerase adds new nucleotides to the growing strand. If the last nucleotide added is mismatched, then the entire replication holoenzyme backs up, removes the last incorrect base, and attempts to add the correct base again. The enzyme is "3' to 5'" because it scans in the opposite direction of DNA replication, which we learned must always be 5' to 3'. The mechanism of the proof-reading system offers an explanation as to why DNA replication must occur in this direction.
Keeping in mind the chemical mechanism we learned for the addition of
nucleotides to the growing DNA strand, imagine what happens when the proof-
reading system removes an incorrectly paired base. The exonuclease removes the
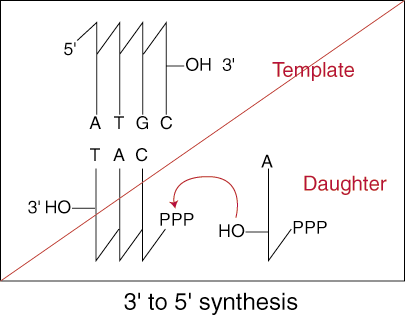
base by cleaving the phosphodiester bond that had just been formed. In 5' to 3'
synthesis, this leaves the 3' -OH still attached to the terminal end of the
growing strand ready to attack another nucleotide.
After DNA has been completely replicated, the daughter strand is often not a perfect copy of the parent strand it came from. Mutations during replication and damage after replication make it necessary for there to be a repair system to fix any errors in newly synthesized DNA. There are three main sources of damage to DNA.
These different sources of damage lead to different categories of DNA damage. The damage that is caused by water attack can lead to unnatural bases. Chemical and radiation damage leads to the formation of bulky adducts to, or breaks in, the growing DNA strand. In the previous section we discussed the 3' to 5' proof-reading exonuclease that is responsible for fixing mismatches. Because it is not a perfect system, it can miss mismatched bases. As a result, a third category of DNA damage is mismatched bases.
Please wait while we process your payment





