Please wait while we process your payment
If you don't see it, please check your spam folder. Sometimes it can end up there.
If you don't see it, please check your spam folder. Sometimes it can end up there.
Please wait while we process your payment
Get instant, ad-free access to our grade-boosting study tools with a 7-day free trial!
Learn more



Create Account
This site is protected by reCAPTCHA and the Google Privacy Policy and Terms of Service apply.
Log into your PLUS account
Create Account
Select Plan
Payment Info
Start 7-Day Free Trial!
Select Your Plan
Monthly
$5.99
/month + tax
Annual
$29.99
/year + taxAnnual
2-49 accounts
$22.49/year + tax
50-99 accounts
$20.99/year + tax
Select Quantity
Price per seat
$29.99 $--.--
Subtotal
$-.--
Want 100 or more? Request a customized plan
Monthly
$5.99
/month + taxYou could save over 50%
by choosing an Annual Plan!

Annual
$29.99
/year + taxSAVE OVER 50%
compared to the monthly price!
| Focused-studying | ||
| PLUS Study Tools | ||
| AP® Test Prep PLUS | ||
| My PLUS Activity | ||
Annual
$22.49/month + tax
Save 25%
on 2-49 accounts
Annual
$20.99/month + tax
Save 30%
on 50-99 accounts
| Focused-studying | ||
| PLUS Study Tools | ||
| AP® Test Prep PLUS | ||
| My PLUS Activity | ||
Testimonials from SparkNotes Customers
No Fear provides access to Shakespeare for students who normally couldn’t (or wouldn’t) read his plays. It’s also a very useful tool when trying to explain Shakespeare’s wordplay!
Erika M.
I tutor high school students in a variety of subjects. Having access to the literature translations helps me to stay informed about the various assignments. Your summaries and translations are invaluable.
Kathy B.
Teaching Shakespeare to today's generation can be challenging. No Fear helps a ton with understanding the crux of the text.
Kay H.
Testimonials from SparkNotes Customers
No Fear provides access to Shakespeare for students who normally couldn’t (or wouldn’t) read his plays. It’s also a very useful tool when trying to explain Shakespeare’s wordplay!
Erika M.
I tutor high school students in a variety of subjects. Having access to the literature translations helps me to stay informed about the various assignments. Your summaries and translations are invaluable.
Kathy B.
Teaching Shakespeare to today's generation can be challenging. No Fear helps a ton with understanding the crux of the text.
Kay H.
Create Account
Select Plan
Payment Info
Start 7-Day Free Trial!
Payment Information
You will only be charged after the completion of the 7-day free trial.
If you cancel your account before the free trial is over, you will not be charged.
You will only be charged after the completion of the 7-day free trial. If you cancel your account before the free trial is over, you will not be charged.
Order Summary
Annual
7-day Free Trial
SparkNotes PLUS
$29.99 / year
Annual
Quantity
51
PLUS Group Discount
$29.99 $29.99 / seat
Tax
$0.00
SPARK25
-$1.25
25% Off
Total billed on Nov 7, 2024 after 7-day free trail
$29.99
Total billed
$0.00
Due Today
$0.00
Promo code
This is not a valid promo code
Card Details
By placing your order you agree to our terms of service and privacy policy.
By saving your payment information you allow SparkNotes to charge you for future payments in accordance with their terms.
Powered by stripe
Legal
Google pay.......



Thank You!
Your group members can use the joining link below to redeem their membership. They will be prompted to log into an existing account or to create a new account. All members under 16 will be required to obtain a parent's consent sent via link in an email.Your Child’s Free Trial Starts Now!
Thank you for completing the sign-up process. Your child’s SparkNotes PLUS login credentials are [email] and the associated password. If you have any questions, please visit our help center.Your Free Trial Starts Now!
Please wait while we process your payment

Sorry, you must enter a valid email address
By entering an email, you agree to our privacy policy.
Please wait while we process your payment

Sorry, you must enter a valid email address
By entering an email, you agree to our privacy policy.
Please wait while we process your payment

Your PLUS subscription has expired
Please wait while we process your payment
Please wait while we process your payment
Month
Day
Year
Please read our terms and privacy policy
Please wait while we process your payment

Transfer RNA
In The Genetic Code, we explained how each codon in messenger RNA (mRNA) codes for a specific amino acid, and that in the process of translation the mRNA brings the amino acids together to form proteins. That explanation is correct, but it is also simplified, and overlooks a crucial component of the translation process. That component is transfer RNA (tRNA), which acts as a kind of link between the information encoded in the mRNA and the amino acids. If the mRNA is a code, then the tRNA is the key that interprets that code into physical proteins.
This section will describe the structure of tRNA and describe how tRNA can "carry" amino acids; knowledge of these aspects of tRNA will be vital for understanding the actual process of proteins synthesis covered next section.
Transfer RNA molecules vary in length between 60 and 95 nucleotides, with the
majority measuring about 75 nucleotides (much smaller than the normal mRNA
strand). Regions of self-complementarity within tRNA creates a cloverleaf-
shaped structure.
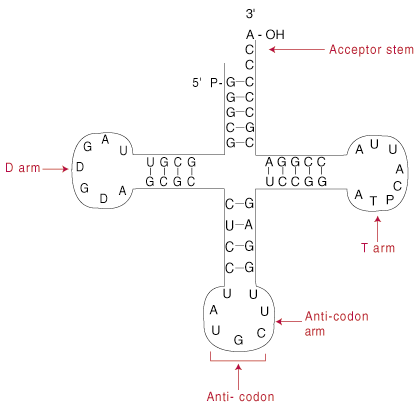
The cloverleaf structure shown above is actually a two dimensional simplification of the actual tRNA structure. The cloverleaf is therefore called a secondary structure. In reality, the cloverleaf folds further into a tertiary structure, a sort of vague L-shape. At one end of the L lies the anticodon; at the other is the acceptor stem. The L-shaped structure simply amplifies the two active ends of tRNA: the anticodon and the acceptor stem.
The structure of the anticodon of tRNA helps to explain the degeneracy of the genetic code. Previously, in the SparkNote on the Genetic Code, we saw that more than one codon could specify a particular amino acid. However, now we know that tRNA acts as a go between for the codons of mRNA and amino acids. Each tRNA binds to a specific amino acid, but the anticodons of some tRNA molecules can bind to two or three different codons.
The flexibility of some anticodons is the result of the fact that the 3' end of the anticodon is more spatially confined than the 5' end. As a result, the 5' end of the anticodon is free to hydrogen bond with several base groups located at the 3' position of the codon. This idea is called the wobble hypothesis and has been confirmed by X-ray studies that show that while the 3' and middle positions are held tightly in a specific orientation by stacking interactions, the 5' position is not. The 5' position is called the wobble position because it can move around to allow for its pairing with different bases.
Please wait while we process your payment





